|
|
|
|||||||||||||||||||||||
|
Tools for Safety AssessmentIdentification and monitoring of Escherichia coli K-12 safety strainsPeter Kuhnert and Joachim Frey Institute of Veterinary-Bacteriology, University of Bern, Länggass-Str. 122, CH-3012 Bern Agency for Biosafety Research and Assessment of Technology Impacts of the Swiss Priority Programme Biotechnology Scope of the bulletinThe bulletin 1/96 appears in the series "Tools for safety assessment". The reader is offered a summary on special subjects with relevance for safety assessment of biotechnology applications. The series would like to meet the need for a more intensive transfer of "first hand" informations in a structured and issue focused synthesis of available knowledge. The bulletin 1/96 has been assembled in co-operation with Dr. P. Kuhnert and Dr. J. Frey of the Institute of Veterinary-Bacteriology, University of Bern. The collaboration was initiated because an unmistakable basis for decision-making in the categorisation of E. coli strains was not available. The research project financed by the Swiss Priority Programme Biotechnology yielded molecular tools which allow an unambiguous classification of E. coli strains. The report which is based on the paper in reference 4, gives a summary of the theoretical background and methodical details. Table of Contents1 Introduction 2 Test system for the identification of Escherichia coli K-12 safety
strains 3 References 1. IntroductionBacterial safety strains are requested for many biotechnological processes, in particular for those involving expression of recombinant genes. Such strains must be non-pathogenic for human, animal and plant. In addition they should exhibit an impaired survival under non-laboratory conditions. The best known bacterial safety strains fulfilling these criteria are the Escherichia coli K-12 derived strains. These strains show the following advantages: i) they represent one of the genetically best understood living organisms; ii) they are classified in all major national and international safety guidelines as biologically safe vehicles for the propagation of a broad range of efficient gene cloning and expression vectors; iii) they are easily modified by many genetic methods, and vi) they exist as a large variety of genetically and phenotypically different mutant variants selected for many specialized scientific and technological applications. While very little is known about the pathogenicity of the wild-type E. coli K-12 strain, which was isolated from a convalescent diphteria patient at Stanford University in 1922, derivatives of E. coli K-12 which were the first time reported in 1944 (ref. 3) seem to be entirely apathogenic. Derivatives of E. coli K-12 were obtained through intensive mutagenesis and extended growth for many passages under laboratory conditions which probably resulted in the loss of all currently known virulence genes (Fig. 1). Moreover, E. coli K-12 derivatives are rough, lacking the 0-antigen which is part of the lipopolysaccharide (Fig. 1). No case of disease, caused by E. coli K-12 derivatives has ever been reported since 1944 in spite of very frequent use of these strains in teaching and research. It was also shown that K-12 strains are unable to colonize the human gut (ref. 8). The K-12 lineage is therefore considered to be a prototype of safe and non-pathogenic bacterial strains. No virulence genes have been found so far in E. coli K-12 derivatives (ref. 7) in contrast to pathogenic E. coli strains which possess normally several chromosomal or plasmid borne virulence genes, which are often located as gene clusters on "pathogenicity islands" (Fig. 1) (ref. 2; 6). This confirms the non-pathogenic phenotype of K-12 derivatives. The absence of virulence genes gives K-12 derivatives further the advantage that no sudden reversion of a mutated gene which might result in the reemergence of pathogenic revertants could occur. 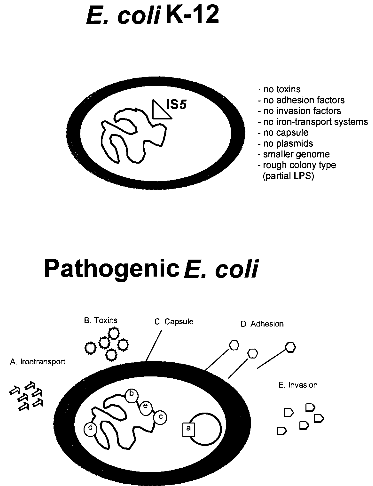
Fig. 1. Comparison of an E. coli K-12 derivative with a pathogenic E. coli strain with regard to virulence attributes and their genes (ref. 7). Shown are two E. coli bacteria and their genomes. E. coli K-12 derivatives do not contain virulence genes. A specific insertion sequence IS5 is contained in the genes encoding the lipopolysaccharide (LPS) pathway genes, rendering these strains rough colony type. The pathogenic E. coli possesses virulence genes a-e encoding the variouse virulence attributes A-E. 1.1. Identification of Escherichia coli K-12The broad range of genetic and phenotypic varieties deduced from K-1 2 and other E. coli strains cause major problems in identifying their correct origins. Entire pedigrees would therefore be required for clear identification of strains. Complete description of bacterial lineages, however, are only available for a limited number of ancestral K-12 strains (ref. 1). This often causes severe problems in the interpretation of experimental data and in particular in biosafety assessments. A particular difficulty is the lack of a common character to E. coli K-12 derivatives which would enable to differentiate them easily from other, apparently very similar E. coli strains. Hence, an accurate and rapid method to discriminate E. coli K-12 from other E. coli strains is needed as a standard tool in biological safety procedures. 1.2. Common genetic marker for E. coli K-1 2 derivativesE. coli K-12 strains are rough, apparently lacking the 0-antigen which is part of the lipopolysaccharide (LPS) and which is encoded by the rfb-gene cluster. Liu and Reeves (ref. 5) recently showed that the lack of 0-antigen in some K-12 derivative strains is due to a mutation (rfb-50) within the rfb-cluster which inactivates the rhamnose transferase, a key enzyme in the 0-antigen biosynthesis. The rfb-50 mutation is characterized by an IS5 insertion that is located within the last gene (orf264 also named orf11 or orf5) of the rfb-cluster which encodes the rhamnose transferase (Fig.2) (ref. 5,9). This mutation was shown to be specific to K-12 derivatives. It can be identified by an appropriately designed ploymerase chain reaction (PCR) (ref. 4). The analysis of 37 K-12 derivatives including strains of all the main branches of the pedigree as well as newly emerged K-12 derivatives showed that they all seem to carry the rfb-50 mutation (ref. 4). In contrast over 70 other E. coli or closely related strains (Shigella and Salmonella) isolated from human, animal, water and soil were devoid of IS5 in the rfb gene. The herein described PCR reaction is therefore considered to be a reliable method for the rapid and accurate identification of E. coli K-12 derivatives (ref. 4). 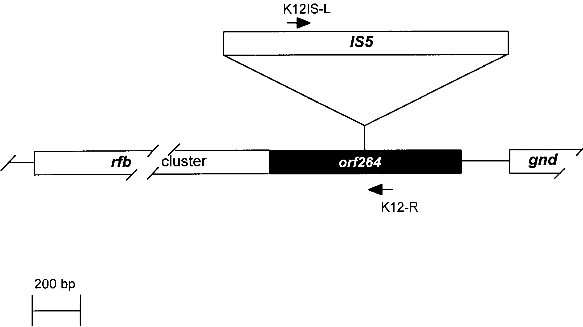
Fig. 2. Map of the rfb-cluster and region analyzed in this study. The rfb-cluster is located 5' to the 6-phosphogluconate dehydrogenase gene (gnd). Boxes represent genes. The entire rfb-cluster extends over more than 10 kb and contains 11 genes. The gene located at the 3' end of the cluster, orf264 (shaded box), encodes the rhamnose transferase. It harbors an IS5 sequence in most K-12 derivative strains specifying the rfb-50 mutation. Arrows indicate the location of primers used for PCR. 2. Test system for the identification of Escherichia coli KA 2 safety strains2.1. Sample preparation for PCRA) By lysate preparation: E. coli strains are grown over night on LB-plates containing, per liter: Three to five colonies are lysed in 450 ml lysis-buffer (100 mM Tris-HCI
pH 8.5, 0.05% Tween20 and 240 µg/ml proteinase K). B) Directly: Alternatively, take 1 µl bacterial liquid culture or few bacteria from a colony taken by a toothpick from a plate, and place them directly in the premix containing Taq-polymerase. 2.2. Oligonucleotide primers
Primer synthesis was done at: Microsynth, Schützenstr.15, CH-9436, Balgach The primer pair K121S-L and K12-R is specific to K-12 and will amplify a 970 bp fragment only in Escherichia coli K-12. The primer pair ECPAL-L and ECPAL-R is specific for all Escherichia coli and some related enteric bacteria and will amplify a control fragment of 280 bp from the exeC gene (PAL protein) (ref. 4). 2.3. Premix for PCR assay1 ml: 2.4. PCR reactionPrepare in a MicroAmpTM (Perkin Elmer Cetus, Norwalk, CT, USA) tube
50 ml premix and add. 2.5. CyclingPlace PCR reaction tube in a PE9600 (Perkin Elmer Cetus, Norwalk, CT,
USA) or equivalent cycling machine and run the following program: 2.6. AnalysisLoad 5 µl of the cycled PCR reaction on a 1% agarose gel containing 500 ng/ml ethidium bromide. Run at 110V for 2h or according to other specifications in TBE-buffer (1 l: 5.4 g Tris, 0.4 g EDTA, 2.75 g Boric acid). 2.7. Identification of E. coli K-12 derivativesE. coli K-12 will produce a specific 970 bp band in addition to the 280 bp control band. If only the 280 bp control band shows up, the strain investigated is not a K-12 derivative. If no band at all shows up, either the reaction was not working properly or the strain investigated is not E. coli or a related species. 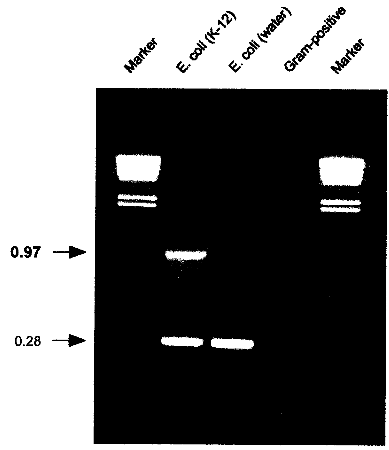
Fig. 3: PCR results showing the K-12 specific band at 0.97 kb resulting from amplification with K121S-L and K12-1R. A natural isolate of E. coli from water does not show the specific band. Control amplifications for E. coli and related enteric species show a band at 0.28 kb resulting from amplification with primers ECPAL-L and ECIDAL-R. Lower bands represent primer-dimer artifacts. A Gram-positive control shows no amplification product at all. Marker: HindIll digested λ DNA with the sizes 23.119.416.614.412.312.010.6 kb. 2.8. ApplicationThe herein described specific PCR assay for the identification of E. coli K-12 derivatives constitutes a useful method in research and quality control for laboratories involved in recombinant genetechnology. It is also a rapid and convenient method to monitor the presence of K-12 derivatives used in biotechnological production processes, thereby providing a basic instrument to contribute to the safety of such processes. 3. References1 . Bachmann, B.J. 1987. Derivations and genotypes of some mutant derivatives of Escherichia coli K-12, In p. 1190-1219. Neidhardt, F.C. (ed.) Escherichia coli and Salmonella typhimurium: cellular and molecular biology. American Society for Microbiology, Washington, D.C. 2. Blum, G., M. Ott, A. Lischewski, A. Ritter, H. Imrich, H. Tschaepe and J. Hacker. 1994. Excision of large DNA regions termed pathogenicity islands from tRNA- specific loci in the chromosome of an Escherichia coli wild-type pathogen. Inf.lmmun. 62..606-614 3. Gray, C.H. and E.L. Tatum. 1944. X-ray induced growth factor requirements in bacteria. Proc. Natl.Acad.Sci. USA 30:404-410. 4. Kuhnert, P., J. Nicolet and J. Frey. 1995. Rapid and accurate identification of Escherichia coli K-12 strains. Appl.Environ. Microbiol. 61:4135-4139. 5. Liu, D. and P.R. Reeves. 1994. Escherichia coli K12 regains its 0 antigen. Microbiology 140:49-57. 6. McDaniel, T.K., K.G. Jarvis, M.S. Donnenberg and J.B. Kaper . 1995. A genetic locus of enterocyte effacement conserved among diverse enterobacterial pathogens. Proc.Natl.Acad.Sci. USA 92:1664-1668. 7. M0hidorfer, 1. and J. Hacker. 1994. Genetic aspects of Escherichia coli virulence. Microbial Pathogenesis 16:171-181. 8. Smith, H.K 1975. Survival of orally administered E. coli K12 in alimentary tract of human. Nature 255:500-502. 9. Stevenson, G., B. Neal, D. Liu, M. Hobbs, N.H. Packer, M. Batley, J.W. Redmond, L. Lindquist and P.R. Reeves. 1994. Structure of the 0 antigen of Escherichia coli K-12 and the sequence of its rfb, gene cluster. J.13acteriol. 176:4144-4156.
|
||||||||||||||||||||||||
|
|




 Produkten.
Produkten.