|
Biochip technology
combines two in one: multiparametric molecular analysis and
quick identification of genomic material in biological samples.
In this article JoséRemacle and Stèphanie Warnon from the
University of Namur (FUNDP, Laboratoire de Biochimie Cellulaire,
Belgium) introduce this promising technology, show its possible
applications and report on current research activities.
General introduction
to biochips
DNA microarrays
or biochips are formed by grafting multiple capture probes
onto a surface. The word "biochip" derives from the computer
term "chip". Although silicon surfaces bearing printed circuits
can be used for DNA binding, the term biochip is now broadly
used to describe all surfaces bearing microscopic spots, each
one being formed by specific capture probes. The capture probes
are chosen to complement the target sequence to be detected.
Each capture probe will bind to its corresponding target sequence.
The purpose of the chips is to detect many genes present in
a sample in one assay rather than performing individual gene
assays as is the practice e.g. in so-called multiwells, plates
with 96 wells, where the reactions take place. The huge amount
of information coming from the genome sequence and other research
genome programs cannot be utilised to the full without the
availability of methods such as biochips which enable these
genes or specific DNA sequences to be detected in biological
samples. DNA microarray technology is an example of the enormous
efforts undertaken in the genomic field in the last few years
Many applications
and technologies adapted to these applications use biochips.
One of the most common applications is in the determination
of gene expression for a given tissue. This is useful in correlating
a change in gene expression with pathology or with cell response
to drugs. These applications have been well developed by companies
like Affymetrix (www.affymetrix.com) or Incyte
(www.incyte.com) using
high density chips bearing either small or long capture probes.
Researchers and pharmaceutical companies chips searching for
new drug targets have displayed most interest in the potential
of such chips.
Chips can also
be utilised in the determination of Single Nucleotide Polymorphism
(SNP) allowing one to discriminate between sequences differing
by a single nucleotide. Using small capture probes (Affymetrix)
and changing the stringency of hybridisation using electric
repulsion (Nanogen http://www.nanogen.com) we have succeeded
in doing this. However the test is far from being quantitative
and sensitivity is very poor. The determination of SNP is
of relevance when studying genetic mutations which can be
linked to genetic disease or in differentiating close species
or homologue genes.
 Top
Top
The proposed strategy
for biochips
We are focusing
our efforts on the developement of biochips as tools for every
day diagnosis so that large numbers of genes or sequences
can be tested in one assay giving the doctor all the information
s/he needs to treat the patient. In practice biochips for
the detection of Staphyloccocus first allows the identification
of Staphylococcus as the source of infection. Secondly
they enable us to establish whether the Staphylococcus
are resistant to methicilin thereby helping to determine the
appropriate antibiotic treatment.
The chips used
are precisely defined products consisting of a fixed and limited
number of carefully chosen capture probes. The choice of the
DNA sequences to be detected is made according to the answer
expected. The chips give the user the information required
but no more. In this way the size of the biochip is maintained
at a minimum and product costs are drastically reduced.
Our experience
in the development of quantitative assays for DNA and fixation
were to our advantage in the optimisation of the DNA probe
hybridisation thereby obtaining assays with good sensitivity
and reproducibility.
Indeed, the main
problem encountered in chip technology is the following: with
miniaturisation of the detection spots, there is a concomitant
lowering of the detection signal and an increase in noise
so that the assay sensitivity is also diminished.
 Top
Top
The proposed chip
technology
A general presentation
of the chips is given in figure 1:
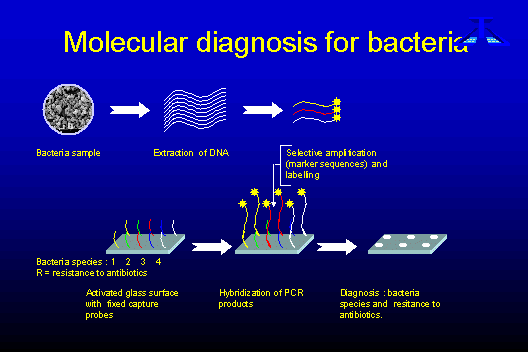
Fixation of the
DNA capture probes
The capture probes
are covalently linked by their 5' end in order to control
both the amount fixed and the length of the sequence available
for hybridisation. The capture probes are single strand and
specially designed for high yield hybridisation. The laboratory
know-how involved here is very important when seeking to obtain
high sensitivity.
Spotting
 We developed a specially devised robot to lay
the capture probes on a glass slide. The robot was developed
in collaboration with WOW (www.wowcompany.com), a company
which has all the knowledge to construct further robots for
production purposes (figure 2). The needles used for a standard
array have a diameter of 0.25 mm (figure 3). With this resolution
a microarray of 1cm2 can hold 400 spots. Variations
are possible since the resolution of the robot is 5 micrometers.
This technology can easily be adapted to obtain a chip with
4,000 spots of 150 micrometers in diameter. The advantages
of the robot are its flexibility, its high output and the
automation of the biochips production process. We developed a specially devised robot to lay
the capture probes on a glass slide. The robot was developed
in collaboration with WOW (www.wowcompany.com), a company
which has all the knowledge to construct further robots for
production purposes (figure 2). The needles used for a standard
array have a diameter of 0.25 mm (figure 3). With this resolution
a microarray of 1cm2 can hold 400 spots. Variations
are possible since the resolution of the robot is 5 micrometers.
This technology can easily be adapted to obtain a chip with
4,000 spots of 150 micrometers in diameter. The advantages
of the robot are its flexibility, its high output and the
automation of the biochips production process.
 Top
Top
Detection
Detection is commonly
performed by fluorescence after incorporation of one fluorochrome
in the target sequences during duplication or amplification.
A double labelling system is useful when comparing two samples.
However, the fluorescent scanner is not an easy machine to
use and is too expensive for clinical laboratories.
We sought to develop
a detection method which could be adapted for routine laboratory
use. We developed a new process which permits a darkening
of the spots testing positive for DNA binding. These spots
can be analysed with a colorimetric scanner with a resolution
of 10 micrometers which is sufficient to quantify the spots.
Analysis software which recognises and identifies the spots
was developed. It also quantifies the labelling and perfoms
a statistical analysis of the data. The scanner is available
together with the software and the computer. This technology
is covered by a filed patent (EU 99870106.4). The main advantage
of this detection system is its simplicity and very low cost.
 Top
Top
The EU Project
"New technology
in food sciences encounter a multiplicity of recently released
gmo's" (gmo chips)"
This year sees
our participation in a new gmo chips project with six other
partners. (www.bats.ch/gmochips/partners/).
The project aims to develop biochips capable of detecting
genetically modified organisms (gmo's) in food. Reliable detection
methods are essential for the labelling of foodstuffs containing
gmo's and for EU border controls.
Over the last years,
there has been a dramatic and continuing increase of the surface
area planted with transgenic crops. The five principal transgenic
foodstuffs are maize, soybean, rapeseed / canola, tomato and
potato. The European Union informs the consumers of the presence
of a transgenic foodstuff by labelling of "substantially equivalent"
ingredients (Food Regulation 258/97 and the Council regulations
1139/98, 49/2000 and 50/2000). The fortuitous presence of
recombinant DNA or modified protein above a defined threshold
of 1 % of ingredients in the foods will lead to unambiguous
food labelling. The development and application of reliable
and quantitative analytical detection methods are thus essential
in order to perform and to control food labelling as well
as for the possible development of "gmo free" production schemes
and to control plant importation at the EU border. Biochip
technology combines the advantages of clear identification
and a multiparametric approach to detect unexpected gmo's
via their specific patterns.
Further information:
|

![]() More details about the GMOchips-project are
available in the Project Introduction
and the Project Summary.
More details about the GMOchips-project are
available in the Project Introduction
and the Project Summary.