|
|
|
||||||||||||||||||||
1.2 Transformation methods and genetic elements introduced into transgenic plantsNumerous methods have been developed that are used to introduce and integrate 'foreign' DNA into plant cells, leading to transformed plant phenotypes. Only those methods used in the transformation of approved agricultural crops will be briefly described below. These are methodologies based on (i) biological vectors, (ii) physical or (iii) chemical methods (occasionally used in combination with electroporation). For a more detailed description of the methodology, including protocols, the reader is referred to Potrykus and Spangenberg (1995). A transformation system should allow for (Niederhauser et al., 1996):
Points one and three are affected primarily by the choice within the first generation of transformants and by long-term selection for a transgenic marker. The fourth feature depends on the choice of the promoter regulating the transcription of the transgene, but possibly also on the presence of targeting sequences that are directing the gene product to certain organelles (e.g. chloroplast transit peptide sequences). The currently used transformation methods do not allow for a precise prediction of the number of copies of the transforming DNA that will be integrated into the plant cell genome. Conventional back-crossing of the transformants with the untransformed phenotype is frequently one technique for reducing the number of copies of the transforming DNA (per haploid genome) to one or a few. Structural integrity of the introduced DNA and the precision with which the boundaries of the sequences which will ultimately integrate into the host genome can be predicted is partially dependent on the choice (if there is one) of the transformation system (see below).
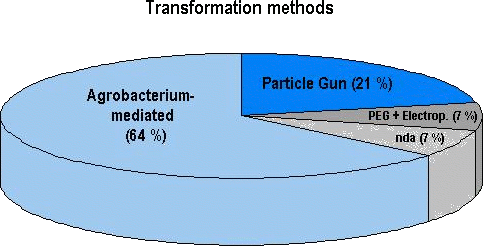
Figure 2: Prevalence of transformation methods used for approved genetically engineered agricultural crops (28 genetically distinct products in total; see later sections). At least 18 out of the 28 products (64 %) have been transformed with Agrobacterium, 6 products with physical methods (e.g. particle gun) and 2 with either a chemical method (e. g. using polyethyleneglycol, PEG) or by electroporation. No data were available (nda) on the transformation system used for 2 of the 28 genetically distinct products. Among the array of genetically engineered plants which have currently
been approved, the transformation method of choice has been the
use of modified plasmids of Agrobacterium (Figure 2). This
is most often a binary vector system derived from Agrobacterium
tumefaciens, where one vector contains the genes to be transferred
and the other harbours genes (vir genes, not transferred)
encoding the necessary functions for transfer to occur (McBride
and Summerfelt, 1990). The system is based on a 'disarmed' Ti-plasmid
with genes responsible for the crown gall disease being removed.
The foreign DNA is confined by the right and left border Other transformation methods are based on physical and chemical principles. According to one method, DNA fragments are bound to the surface of minute metal particles and shot into plant cells using specially developed devices. The chemical methods make use of polyethyleneglycol (PEG) or CaCl2 to facilitate the entrance of foreign DNA through the plant cell wall. Electroporation represents another transformation method. Plants transformed using electroporation, chemical or physical methods generally carry copies of the entire DNA fragments presented. These plants may thus contain copies of the antibiotic resistance genes used for the propagation of the respective constructs in bacteria, if such has not been prevented by removing the respective genes through restriction enzymes prior to transformation. Frequently, some sections of the presented plasmid sequences are not transferred using these methods. Therefore, the boundaries of the transferred DNA will be predicted with less precision using these methods than with Agrobacterium-mediated transformation methods. 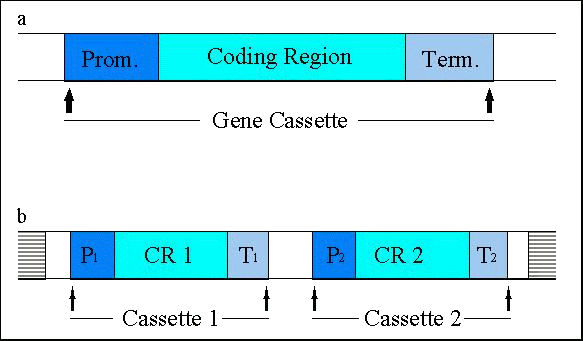
Figure 3: (a) Schematic representation of gene cassettes, consisting of a promoter (P), a structural gene ('coding region') and a terminator (T); (b) frequently, two (or more) cassettes are transferred together and integrated into the host genome (horizontally bars) at one or several sites. Enzymes are the products of the majority of transgenes introduced into the currently approved genetically engineered agricultural crops. The expression of these enzymes has conferred novel traits to the respective plants. Proteins without an enzymatic activity, such as viral coat proteins or the Bt-toxin (-endotoxin from Bacillus thuringiensis), or antisense constructs have also been expressed. Efficient expression of structural genes is assured only when they are controlled by plant-derived promoters or by other promoters that are active in plant cells such as the cauliflower mosaic virus 35S promoter. Terminator sequences also have to originate from plant sources or from plant pests such as the cauliflower mosaic virus or Agrobacterium. Direct selection for the many of the actual trait genes (e.g. those conferring delayed fruit ripening) is not possible. Therefore selectable marker genes, such as genes allowing growth in the presence of antibiotics or herbicides, are often co-transformed with the actual trait gene(s) together with appropriate regulatory sequences (Figure 3b). The final number of 'foreign' gene cassettes that are present in a transgenic crop may be as high as 4 or 5 due to the presence of multiple trait genes and marker gene(s) (Table 3: Approved genetically modified crops in the United States)
|
|||||||||||||||||||||




 antibodies on hand
antibodies on hand 